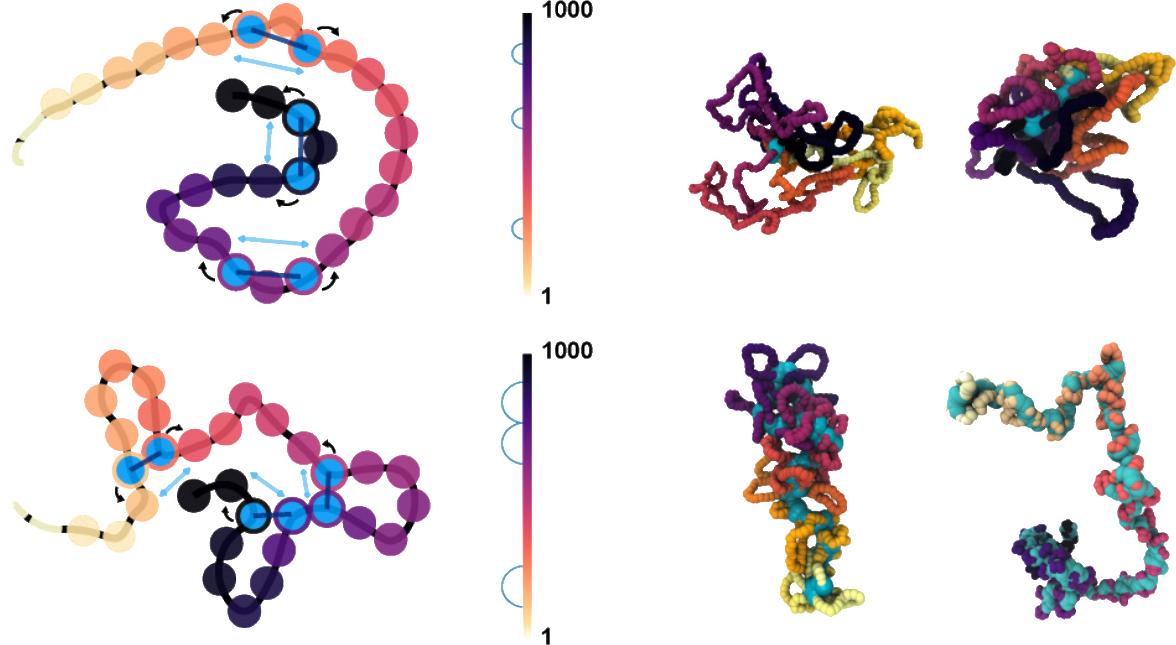
Loop extrusion is one of the main processes shaping chromosome organization across the cell cycle, yet its role in regulating deoxyribonucleic acid (DNA) entanglement and nucleoplasm viscoelasticity remains overlooked. We simulate entangled solutions of linear polymers under the action of generic loop extruding factors (LEFs) with a model that fully accounts for topological constraints and LEF-DNA uncrossability. We discover that extrusion drives the formation of bottlebrushlike structures which significantly lower the entanglement and effective viscosity of the system through an active fluidification mechanism. Interestingly, this fluidification displays an optimum at one LEF every 300–3000 base pairs. In marked contrast with entangled linear chains, the viscosity of extruded chains scales linearly with polymer length, yielding up to 1000-fold fluidification in our system. Our results illuminate how intrachain loop extrusion contributes to actively modulate genome entanglement and viscoelasticity in vivo.
Download “Article preprint” active_fluidification.pdf – Downloaded 357 times – 2 MB
Download a copy of the manuscript (preprint version)